Click the image below to view the full size version of this cover.
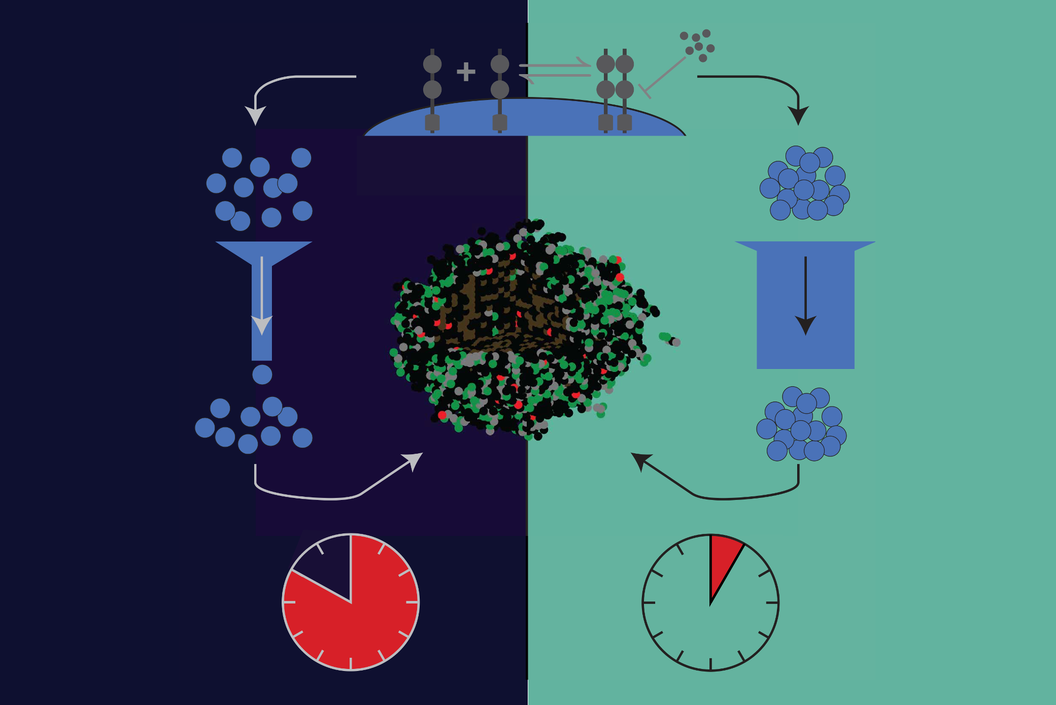
Created by: Daniel Bergman
Issue 214: Agent-based models of biological tissue are computationally expensive. Many models that include molecular dynamics, and intracellular signaling in particular, spend much of their simulation time solving these dynamics. We developed and analyzed an approach to simulating ABMs that aims to reduce this time by coarse-graining these dynamics. This global method for simulating molecular dynamics is 1-2 orders of magnitude faster and produces very similar results to the fine-grained, local method. Learn more about the method, how well it performs, and some future directions in our recent publication here. Check out our blog post for a bit more on how we came up with the method and where we're going.