Mathematical modelling predicts selective sweeps are restricted to early stages of tumour growth
Behind the paper
Selective sweep probabilities in spatially expanding populations
Alexander Stein, Kate Bostock, Ramanarayanan Kizhuttil, Maciej Bak and Robert Noble
Read the paperIn most cancer types, one observes a handful of clonal driver mutations (present in all cells) and, depending on the sampling scheme, one or few subclonal driver mutations (present in only a subset of cells). A natural and fundamental question to ask is when do driver mutations fixate, and when do they remain subclonal due to clonal interference? Because of clinical and experimental constraints, it remains difficult to answer this question directly from sequencing data. However, as we show in our new paper, one can tackle this question through tractable mathematical models and agent-based simulations of tumour growth.
By investigating the timing and frequencies of selective sweeps across different mathematical models, we find that sweeps are rare except during early tumour growth. Once a tumour is in its final growth phase and is more than a cubic millimeter in volume, we predict that even extremely strong drivers are highly unlikely to become clonal and will instead contribute to genetic heterogeneity and possibly to parallel evolution.
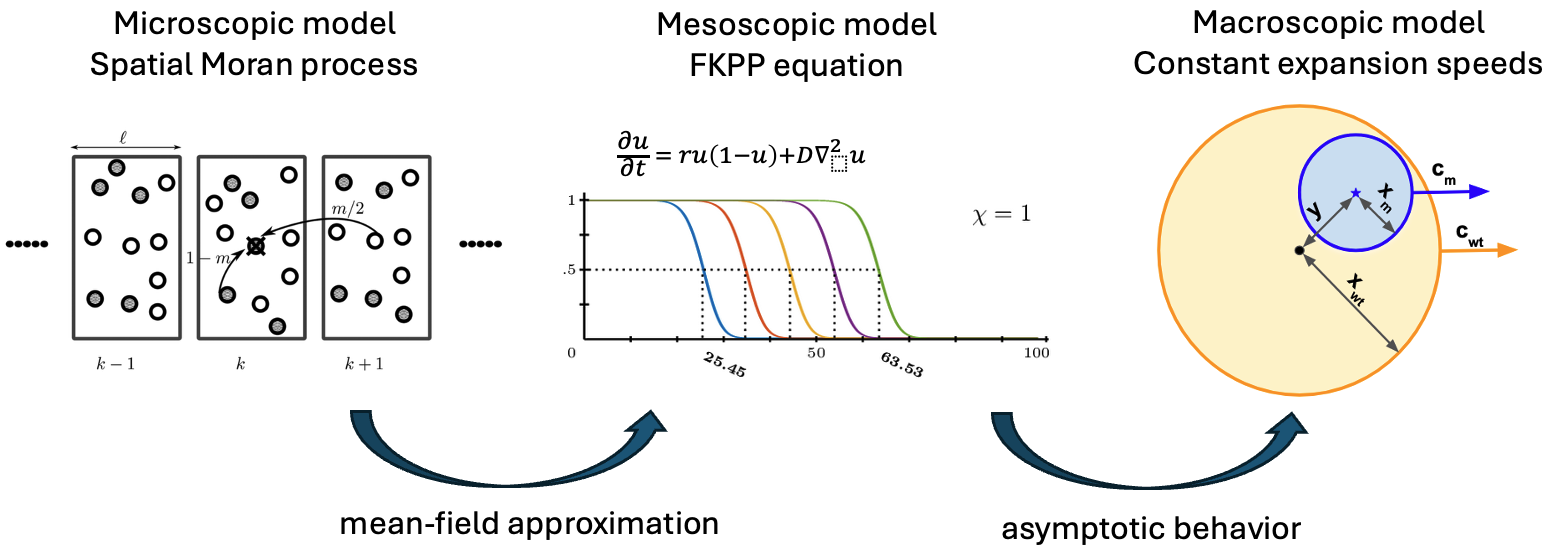
A mathematically tractable model of spatial tumour evolution
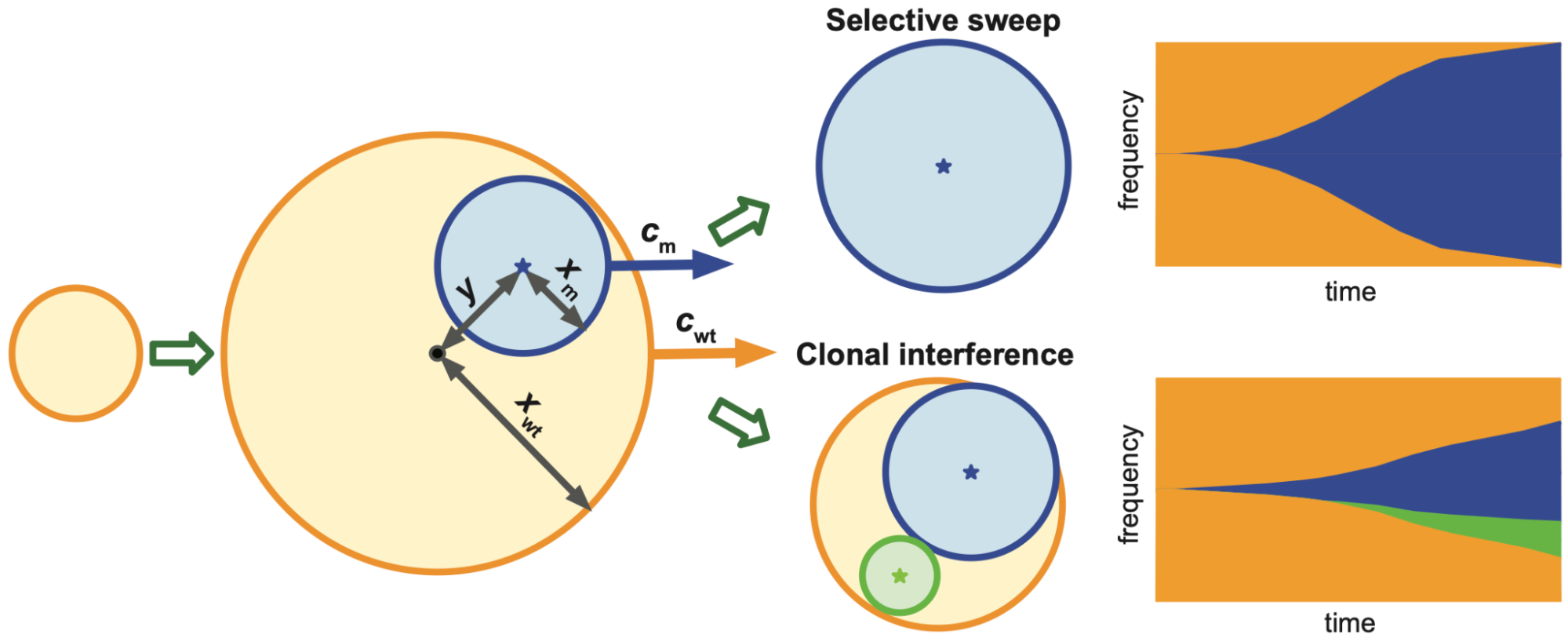
Most models of spatial tumour evolution are either deterministic partial differential equations or stochastic agent-based simulations. In our study, we considered an alternative approach in which the wildtype and mutant populations grow deterministically, but mutations occur stochastically. The model is then an inhomogeneous Poisson process that is relatively easy to analyse and yet yields valuable insights. Combining basic probability theory, integral calculus, and geometry, we derive probability densities for the time when the first mutant occurs, the location of the first mutant and the sweep probability. If the wildtype and the mutant tumours grow at constant speeds, we find a surprisingly simple approximate formula for the sweep probability, namely $$ Pr(sweep) \approx \left(1-\frac{c_{wt}}{c_{m}}\right)^3, $$ where $c_{wt}$ is the propagation speed of the wildtype clone, $c_{m}$ is the propagation speed of the mutant clone (see Figure 1).
Interestingly, we find that the sweep probability is independent of the mutation rate. Intuitively, this can be understood through two opposing effects. First, a higher mutation rate leads to the earlier appearance of the first (sweeping) mutant. As a result, the wildtype population is still smaller, so there are fewer individuals to replace, which increases the sweep probability. Second, a higher mutation rate increases the likelihood that competing mutant clones arise, which decreases the sweep probability. When the propagation speed is constant, these two effects cancel each other out. For exponentially growing populations, we find that the sweep probability slowly increases with increasing mutation rate. This contrasts with constant-size populations, where the first effect does not apply and sweep probabilities are rapidly decreasing with increasing mutation rate.
Investigating successive mutation and random mutation effects
We ran and analysed 68,000 agent-based simulations to validate and extend our results. The first 24,000 simulations demonstrated that our analytic predictions are accurate when mutation effects are fixed (see black triangles vs green curves in Figure 2). In the other 44,000 simulations, we extended our model to allow for successive mutations with random mutation effects. If mutation effects are random, but we allow at most one driver mutation in each cell, we find sweep probabilities similar to those when mutation effects are fixed (black triangles vs red dots in the left panel of Figure 2). If we allow successive driver mutations, sweep probabilities are elevated (cyan square in the left panel of Figure 2). Sweeps also occur later but are still expected to be completed by the time the tumour reaches typical detection sizes (right panel of Figure 2).
This work has started from an idea in 2017. If you are curious about the journey from the conception to the final publication, have a look at this link. We also created some art work that you can find here.
© 2026 - The Mathematical Oncology Blog