Click the image below to view the full size version of this
cover.
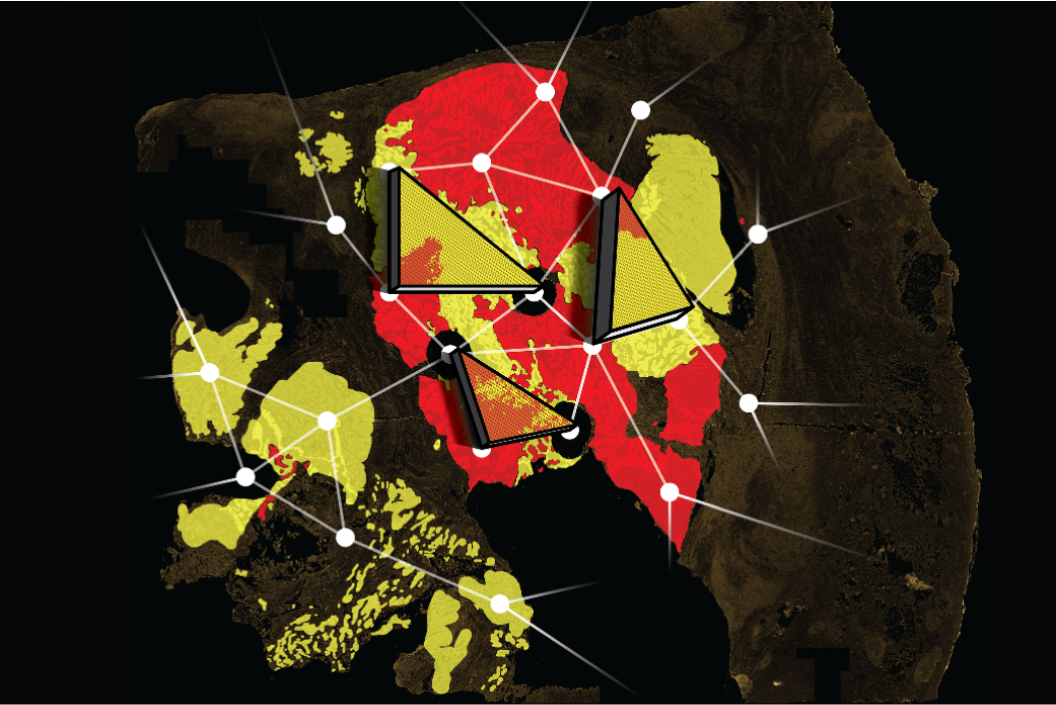
Created by: Magnus Haughey, Ann-Marie Baker, Weini Huang
Issue 257: How do geometrical patterns of tumour sub-populations relate to their underlying dynamics? In our recent study, we set out to tackle the problem of how to quantify tumour sub-clonal dynamics in single time-point human colorectal cancer (CRC) samples using only the information contained within the spatial architecture of different tumour sub-populations. By simulating random walks on the pixels of high-resolution images of human CRC, we exploited the first passage times between the wild-type and mutant KRAS, BRAF, and PIK3CA populations to quantify their observed geometry. Comparing these to similar measurements of simulated spatial tumours, generated using a spatial agent-based model, we then estimated the parameters of our model which best recapitulated the observed sub-population architectures in patient samples. Our results suggested that, despite containing almost no genetic information, it is still possible to extract some information about tumour sub-clonal dynamics by analysing the shape of cell sub-populations.
© 2026 - The Mathematical Oncology Blog