In silico clinical trials: Improving the discovery to approved pipeline
Behind the paper
In silico trials predict that combination strategies for enhancing vesicular stomatitis oncolytic virus are determined by tumor aggressivity
Adrianne L Jenner, Tyler Cassidy, Katia Belaid, Marie-Claude Bourgeois-Daigneaults, Morgan Craig
Read the paperOverview
While mathematical modelling has provided significant advancements in cancer treatment, the speed at which we are moving towards effective all-encompassing treatments is slow in comparison to the growth of this disease1 (Fig. 1). Fortunately, virtual or in silico clinical trials are ideally situated to respond to the needs of the oncology industry2 and assist in more rapid translation of treatments from discovery to approved. Using a mathematical model as a base, in silico clinical trials are designed to capture population variability and determine optimal therapeutic regimes for individuals that may also be robust at the cohort level2. Virtual clinical trials (or in silico ‘twins’) have been used in preclinical research to make ‘go or no go’ decisions3,4. Through applying this technique more readily to models in oncology, we may be able to increase the number of treatments that reach the approved stage by explored more exhaustively the protocols which may result in a robust response at the personal and cohort level. In this paper5, we applied an in silico clinical trial to dual oncolytic virotherapy and predicted dosage protocols that can improve treatment response.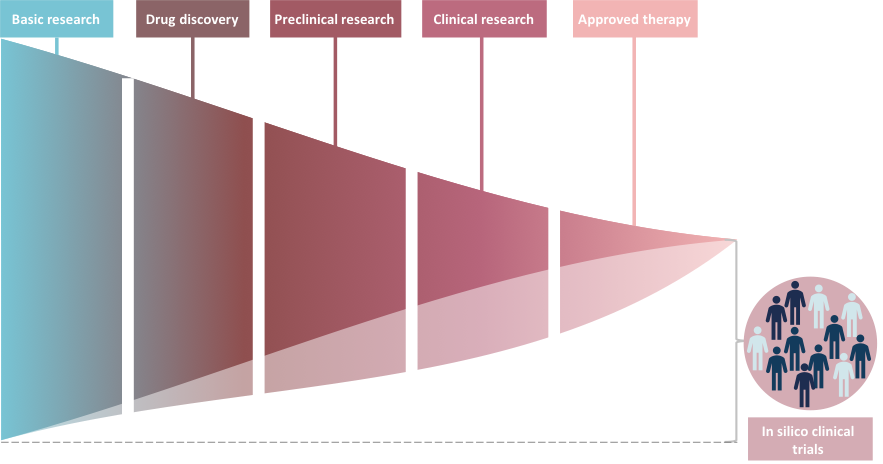
Figure 1: Schematic illustration of the cancer treatment approval process. The number of cancers drugs that achieve approved status are a small proportion of the number originally discovered. In silico clinical trials are a way or generating virtual patients on which to run treatment protocols. It is possible that through applying these techniques we may be able to widen the portion of drugs that make it through to pre-clinical, clinical and even approved therapeutic stage.
Generating an in silico trial for dual OV therapy
Oncolytic viruses (OVs) are viruses genetically designed to infect and replicate within tumour cells6. Through their replication, OVs induce necrosis (cell death) in infected cells, causing a reduction in tumour size6 and instigating an immune response that can be primed towards the tumour6. The synergy between combining OVs has only recently started to gain traction7,8. The principal idea of combination or dual OV therapy is to stimulate alternate paths of antitumour immunity that can act cooperatively to enhance therapy success9. In particular, combining the oncolytic forms of vaccinia (VV) and vesicular stomatitis virus (VSV) has been considered as a potentially effective dual OV therapy as VV is able to enhance VSV’s ability to replicate and spread by dampening down the antiviral result10. To further the pre-clinical investigation of dual OV therapy with VV and VSV, we simulated a virtual (in silico) clinical trial with this therapy using 200 virtual patients (Fig. 2). Each virtual patient was assigned a unique set of parameters, generated to reflect interindividual variability and the heterogeneity in treatment outcomes. To avoid the inclusion of non-realistic virtual individuals, each parameter set was verified as resulting in a tumour growth within two standard deviations of the experimental control tumour growth measurements. With these patients, we then tested a range of alternate schedules to infer realistic dual OV schedules that could have a robust effect clinically.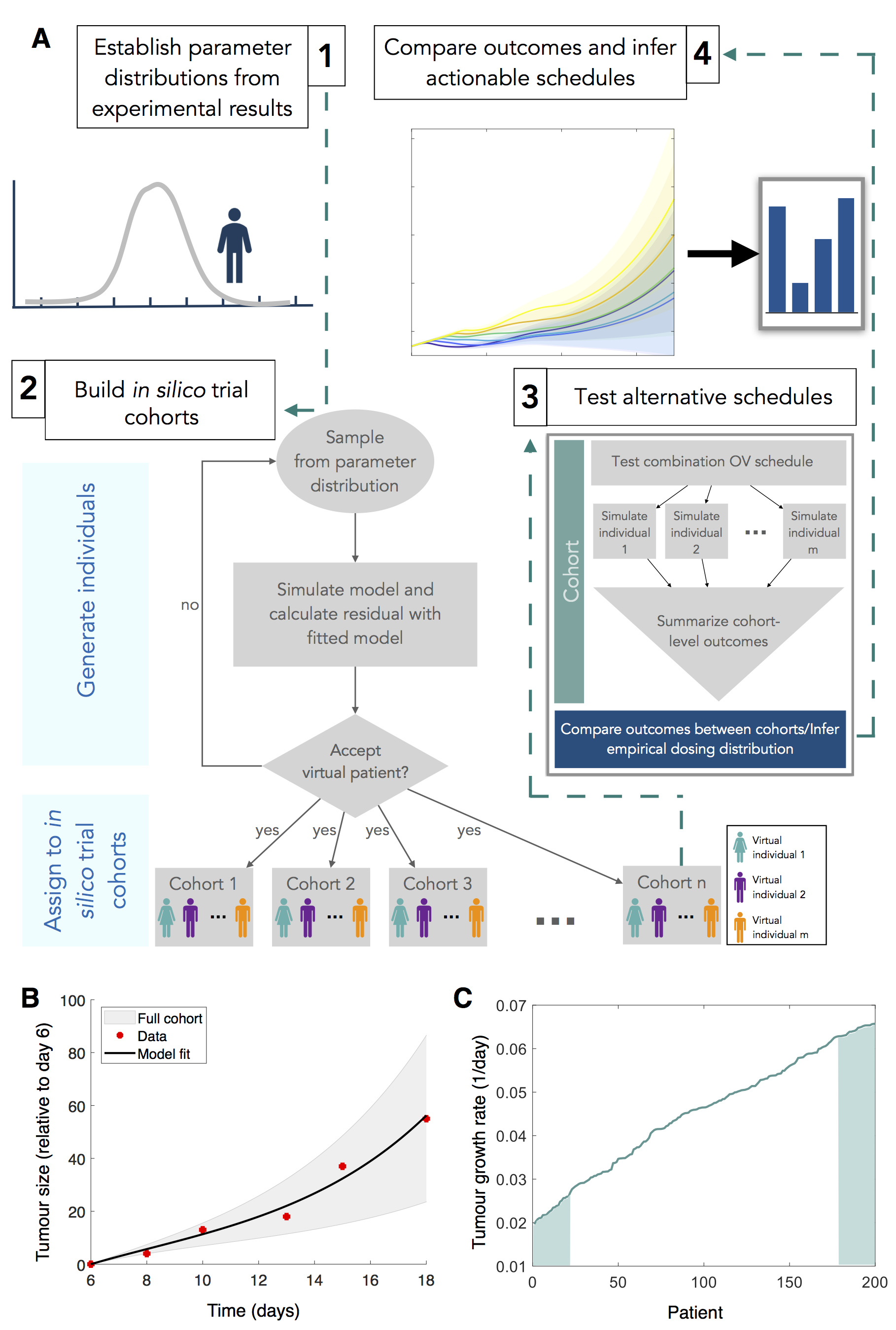
Figure 2: in silico trial strategy for generating virtual patients and predicting optimal clinical schedules. A (1) Parameters are first established by calibrating model predictions to experimental results. A distribution of responses centered at the mean of the experimental data are then used to generate parameter sets representing virtual individuals. (2) To populate the trial, each virtual patient’s tumor growth is simulated to determine whether they are candidates for the trial. Patients whose tumor growth is acceptable (i.e., clinically relevant) are placed into repeated identical cohorts. (3) Alternative treatment schedules are then tested on each cohort by simulating individual virtual patient responses with the mathematical model and summarizing cohort level outcomes. (4) Optimal actionable schedules are then inferred by comparing cohort level and individual outcomes. B Tumour growth over time in the absence of treatment. C Virtual patients were ordered based on intrinsic tumour growth rates (top and bottom 10% denoted by shaded regions).
Optimal protocols are determined by tumour aggressivity
Two large unknowns for dual OV therapy, are the effect of multiplicity of the primary injection and the time lag between the different OV dosages. To assess the impact of these two variables, we determined the optimal number of VV injections (between 1 and 7) and VSV lag (between 1 and 15 days) for each virtual individual (Fig. 3). We then plotted each patient’s optimal protocol against their intrinsic tumour growth rate to determine whether there was any correlation between protocol and tumour aggressivity. We found clear stratification between optimal protocols for slowly growing and aggressively growing tumours: slow growing tumours required 7 VV injections followed by 14-15 day VSV lag and aggressively growing tumours required 1 VV injection followed by a 1 day VSV lag. Interestingly, for a range of low intrinsic growth rates, optimal schedules resulted in a near complete tumour removal; whereas, we found a jump in the number of tumours cells, followed by a linear dependence on the intrinsic growth rate after a critical value of around 0.03/day (roughly equivalent to a doubling time of 23 days). These results suggested that tumour aggressivity could be the determining factor for combination OV scheduling and the outcome of dual VV and VSV OV therapy.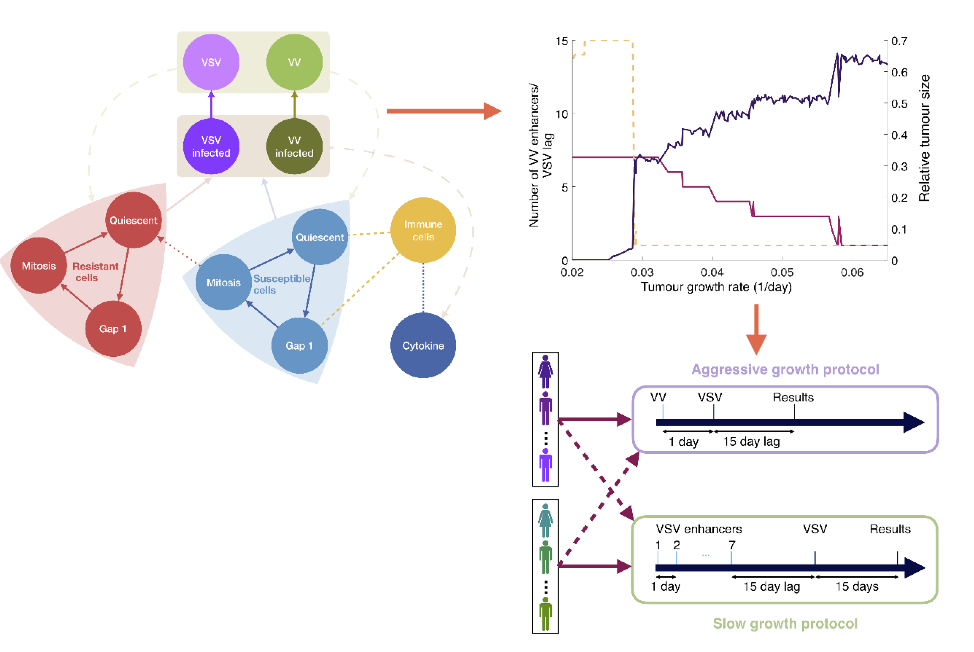
Figure 3: in silico trial predicts that optimal dual OV strategy is determined by tumour aggressivity. The optimal dual OV schedule (number of VV injections and lag between last VV injection and first VSV injection) was determined for each individual virtual patient by simulating the full model (left). In our model, we considered explicit populations of resistant (red) and non-resistance (blue) cycling tumour cells that were infected by either VV or VSV. For each patient, we plotted the optimal number of VV injections (yellow), optimal VSV lag (fuchsia) and relative tumour size 15 days after the last VSV injection (purple) as a function of the patient intrinsic tumour growth. For all but a subset of the least aggressive tumours, individualised protocols called for a VSV lag of 1 day, with fewer than seven enhancers. From this we could deduce the optimal protocol for aggressive and slow growth tumours.
Moving forward
Through using an in silico clinical trial, we recommended a clinically relevant scheduling that can easily be translated from bench to bedside. Broadly speaking, we found that for tumours with low intrinsic growth rates required a larger number of VV injections to be affected, whereas aggressive tumours required fewer VV injections. The latter finding supports the idea of ‘hitting hard, hitting early’ for fast growing tumours. While there are currently no clinical trials combining VV and VSV, other clinical trial results have found a clear distinction in survival based on dosage and the presence of metastases11,12. A high-dose OV protocol on triple negative breast cancer patients with and without metastases performed equivalently, whereas, the low dose OV protocol only performed well for patients without metastases11. As the presence of metastases can be a sign of disease aggressivity, these findings align with our conclusions that the stage and overall aggressiveness of a patient’s cancer is a determinant for dual OV treatment efficacy. Overall, while the drug discovery to approved treatment pipeline is slow, here we put forward a simple strategy that can be used to widen the possible avenues to consider in treatment protocols in a fast efficient manner.References
- Zhang, Z. et al. Overcoming cancer therapeutic bottleneck by drug repurposing. Signal Transduct. Target. Ther. 5, 1–25 (2020).
- Alfonso, S., Jenner, A. L. & Craig, M. Translational approaches to treating dynamical diseases through in silico clinical trials. Chaos An Interdiscip. J. Nonlinear Sci. In review, (2020).
- Allen, R. J., Rieger, T. R. & Musante, C. J. Efficient Generation and Selection of Virtual Populations in Quantitative Systems Pharmacology Models. CPT Pharmacometrics Syst. Pharmacol. 5, 140–146 (2016).
- Wang, H. et al. In silico simulation of a clinical trial with anti-CTLA-4 and anti-PD-L1 immunotherapies in metastatic breast cancer using a systems pharmacology model. R. Soc. open Sci. 6, 190366 (2019).
- Jenner, A. L., Cassidy, T., Belaid, K., Bourgeois-Daigneault, M.-C. & Craig, M. In silico trials predict that combination strategies for enhancing vesicular stomatitis oncolytic virus are determined by tumour aggressivity. J. Immunother. Cancer Accepted, 1–30 (2021).
- Ilett, E. et al. Prime-boost using separate oncolytic viruses in combination with checkpoint blockade improves anti-tumour therapy. Gene Ther. 24, 21–30 (2017).
- Aitken, A. S. et al. Brief Communication; A Heterologous Oncolytic Bacteria-Virus Prime-Boost Approach for Anticancer Vaccination in Mice. J. Immunother. 41, 125–129 (2018).
- Pol, J. et al. Development and applications of oncolytic Maraba virus vaccines. Oncolytic Virotherapy Volume 7, 117–128 (2018).
- Russell, L., Peng, K. W., Russell, S. J. & Diaz, R. M. Oncolytic viruses: priming time for cancer immunotherapy. Springer (2019).
- Le Boeuf, F. et al. Synergistic interaction between oncolytic viruses augments tumor killing. Mol. Ther. 18, 888–895 (2010).
- Heo, J. et al. Randomized dose-finding clinical trial of oncolytic immunotherapeutic vaccinia JX-594 in liver cancer. Nat. Med. 19, 329–336 (2013).
- Zhu, X. et al. The prognostic and predictive potential of Ki-67 in triple-negative breast cancer. Sci. Rep. 10, 1–10 (2020).
© 2026 - The Mathematical Oncology Blog