Mechanistic Learning
The best of both worlds.
A review of mechanistic learning in mathematical oncology
John Metzcar, Catherine R. Jutzeler, Paul Macklin, Alvaro Kohn-Luque, Sarah C. Bruningk
Read the paperWe propose to name such hybrid approaches combining big data and ML with mechanistic modeling "mechanistic learning."To my mind, the only controversial aspect of Benzekry's suggestion is his use of the royal "we" for a single author paper (I joke...).
Mechanistic learning in math oncology
A few years later, John, Catherine, Paul, Alvaro, and Sarah take up the charge and write an interesting and fresh perspective on how to make progress [2]. Below, I will review their paper and comment on why I find it exciting. The brief overview of the history of mathematical modeling in the Introduction is excellent. It is a daunting task to review "what is mathematical modeling?" The authors use the phrase knowledge-driven modeling, and state how constructing these types of models should provide an approximate of our collective, biomedical understanding. The goal is to be precise in writing down the key assumptions, although at times the mathematical descriptions may indeed be "accurate descriptions of our pathetic thinking" [3]. Knowledge-driven modeling stands in contrast to data-driven modeling. The goal here is the extraction of information from existing data. These are the machine learning, deep learning, and other data-hungry AI algorithms. Generalization beyond the particular dataset may be a challenge, and models are vulnerable to overfitting. Post hoc processing is need to unveil the black box's decision-making process.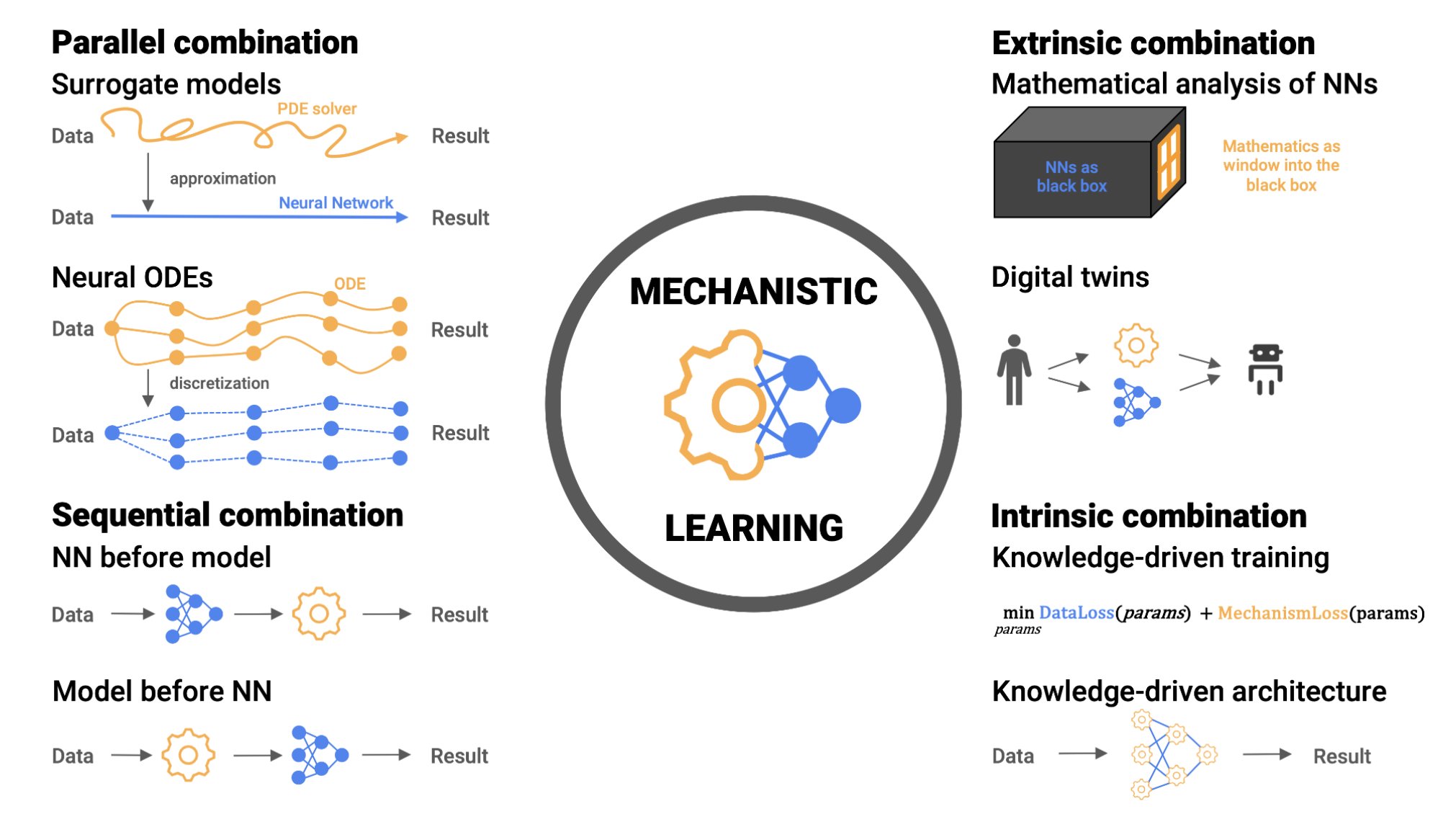
Knowledge + Data = Mechanistic Learning
It's clear that both machine learning and mechanistic models have advantages. Mechanistic models excel at taking sparse datasets that incorporate dynamics (time-dependence) and make remarkable predictions with a few parameters. In contrast, it's not always clear how to incorporate time-dependent information into machine learning methods. There are some methods (recurrent neural networks), but these exceptions prove the rule: they are difficult to train, still data-hungry, and feel like putting a square peg into a round hole (see also: encoding time series as images).Types of Mechanistic Learning
The authors introduce four possibilities of combining knowledge and data-driven methods: 1. Sequential - Knowledge-based and data-driven modeling are applied sequentially building on the preceding results 2. Parallel - Modeling and learning are considered parallel alternatives to complement each other for the same objective 3. Extrinsic - High-level post hoc combinations 4. Intrinsic - Biomedical knowledge is built into the learning approach, either in the architecture or the training phase The first type is arguably the most intuitive. We already typically begin a project by fitting some data with a mechanistic model, resulting in a distribution of patient-specific parameters. What do you do with that list of parameters? One straight-forward option is to run them through a machine learning model, to see if they predict relevant clinical outcomes. In doing this, you have effectively translated the information from time series data into patient-specific parameters, thus resolving the issue that machine-learning models often do not play nice with time-series data. We are trying this approach out in B-precursor ALL. We have Flow Cytometry characterization data for many patients, which gives us information about cell types in each patients. We use this data to train a mechanistic Markov chain model to predict transition rates between cell types. We then train a machine learning model on the Markov parameters. This allows us to determine which parameters are predictive of important outcomes (e.g. minimal residual disease). We were pleased to see the process of mechanistic learning outlined so clearly in the recent review paper by Metzcar and colleagues, which sparked some follow up lines of research in our projects, too. It also works in reverse: data-modeling provides a platform for subsequent mechanistic approaches. Recent work from Heyrim Cho, Russ Rockne et. al. in AML [4] starts with high-dimensional data (single-cell RNA-sequencing) and then the authors reduce the dimensionality of the dataset (e.g. PCA, t-SNE, etc). This reduced space is then used to model flow and transport between differentiation cell states using PDE's acted out in reduced-dimensional space. Importantly, that space is interpretable using pseudo-time analysis.Varia
In the assortment of other reasons I liked the paper is the authors' definition of mathematical modeling.An alternative is to formulate a specific guess on how relevant variables interact between input and output through the formulation of a mathematical model.Mathematicians (mathematical biologists) are proposing their 'guess' of how the world works. This guess of course uses reason, intuition, historical evidence, and most importantly external validation. But the starting point is a reach out into the dark, grasping for the truth. I need not convince the reader that the guessing is a worthy exercise, in itself. Similarly, the authors are firmly in favor of their fellow mathematicians:
It is tempting to suggest that knowledge-driven models are inherently interpretable. Yet, the implementation of chains of relationships can formulate complex inverse problems. Subsequently, post hoc processing through parameter identifiability and sensitivity analyses is key. This can identify previously unknown interactions between system components to generate hypotheses for experimental and clinical validation.In summary, I encourage you to read the review paper. I think it provides several promising strategies to merge two distinct fields that have each earned a solid reputation for progress in math oncology.
References
- Benzekry, S., 2020. Artificial intelligence and mechanistic modeling for clinical decision making in oncology. Clinical Pharmacology & Therapeutics, 108(3), pp.471-486.
- Metzcar, J., Jutzeler, C.R., Macklin, P., Köhn-Luque, A. and Brüningk, S.C., 2024. A review of mechanistic learning in mathematical oncology. Frontiers in Immunology, 15, p.1363144.
- Gunawardena, J., 2014. Models in biology: ‘accurate descriptions of our pathetic thinking’. BMC biology, 12, pp.1-11.
- Cho, H., Ayers, K., DePills, L., Kuo, Y.H., Park, J., Radunskaya, A. and Rockne, R., 2018. Modelling acute myeloid leukaemia in a continuum of differentiation states. Letters in biomathematics, 5(Suppl 1), p.S69.
© 2026 - The Mathematical Oncology Blog