Clonal competition in ageing human haematopoiesis and its emerging dynamic fitness landscape
Behind the paper
The dynamic fitness landscape of ageing haematopoiesis through clonal competition
Nathaniel Mon Père, Francesco Terenzi, Benjamin Werner
Read the paperWe are very happy to talk about our newest preprint on clonal competition as a proposed mechanism for the emerging dynamic fitness landscape of haematopoietic stem cells (HSCs) [1]. This work started with two very exciting papers published back-to-back by Mitchell et al. and Fabre et al. that measured aspects of clonal haematopoiesis (CH) through single cell phylogenies [2] and clonal trajectories respectively [3].
Fabre et al. [3] observed that variant trajectories follow logistic curves. This fits well with the notion of positive selection within a constant HSC population. However, they also noted that this logistic model tended to overestimate clone sizes later in life, leading them to suggest some “deceleration” effect is at play. Looking at the data, we were surprised to find that 20-30% of all variant trajectories were in fact decreasing, with this proportion steadily increasing from a fifth at age 63 to almost half at 88. Many of these trajectories were observed at relatively high allele frequencies, indicating that they must have carried some selective advantage in the past
Mitchell et al. [2] observed balanced (i.e. neutral looking) phylogenetic trees from single-cell HSCs throughout early adulthood, followed by a very sudden transition towards imbalanced trees around the age of 63, suggesting an apparent sudden boom in selection. A curious and dramatic observation.
Before these observations, clonal haematopoiesis was considered the result of rarely occurring HSC variants expanding under strong positive selection. While this perspective provides a simple explanation for the occurrence of CH, it fails to capture all aspects of the data, such as the deceleration of trajectories and the occurrence of negatively selected clones. To reconcile this, we looked for a scenario that could naturally explain these observations together. One possibility was that competition of more-frequently emerging clones dynamically alters the HSC fitness landscape with age. The increasing burden of competition in the system would then reduce the effectiveness of all expanding clones, and eventually cause the least fit clones to become negatively selected and thus switch from growing to shrinking (Figure 1).
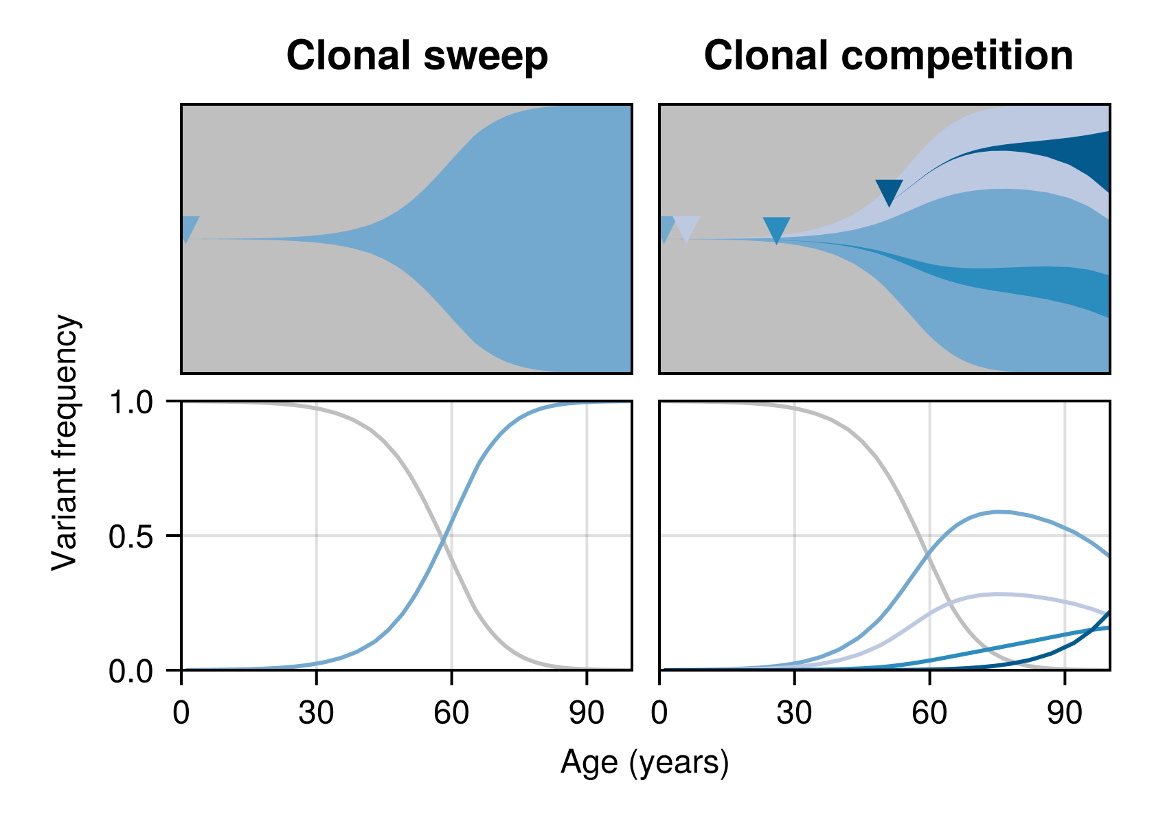
To probe this idea quantitatively, we developed a mathematical model to describe a population of cells in which arbitrary many rivaling clones of varying fitness arise and compete over time. We found the size of individual clones to adhere to a set of stochastic differential equations:
$$\mathrm{d} x_i(t) = x_i(t) \left( s_i - \sum_{\text{all clones}} s_j x_j(t) \right) \mathrm{d} t + B(t) \mathrm{d} W_t$$
with $s_i$ the innate fitness of clone $i$. With novel clones arising as a Poisson process and their innate fitness drawn from a Gamma distribution, we found the model to immediately predict decelerating logistic trajectories and expanded clones that shrink (Figure 2), just as in the Fabre data.
In fact, it furthermore predicted that attempting to infer fitnesses of identical variants in different individuals and at different ages results in a distribution of values (Figure 2), a phenomenon which has been observed experimentally before (for example by van Zeventer et al. [4] amongst others).
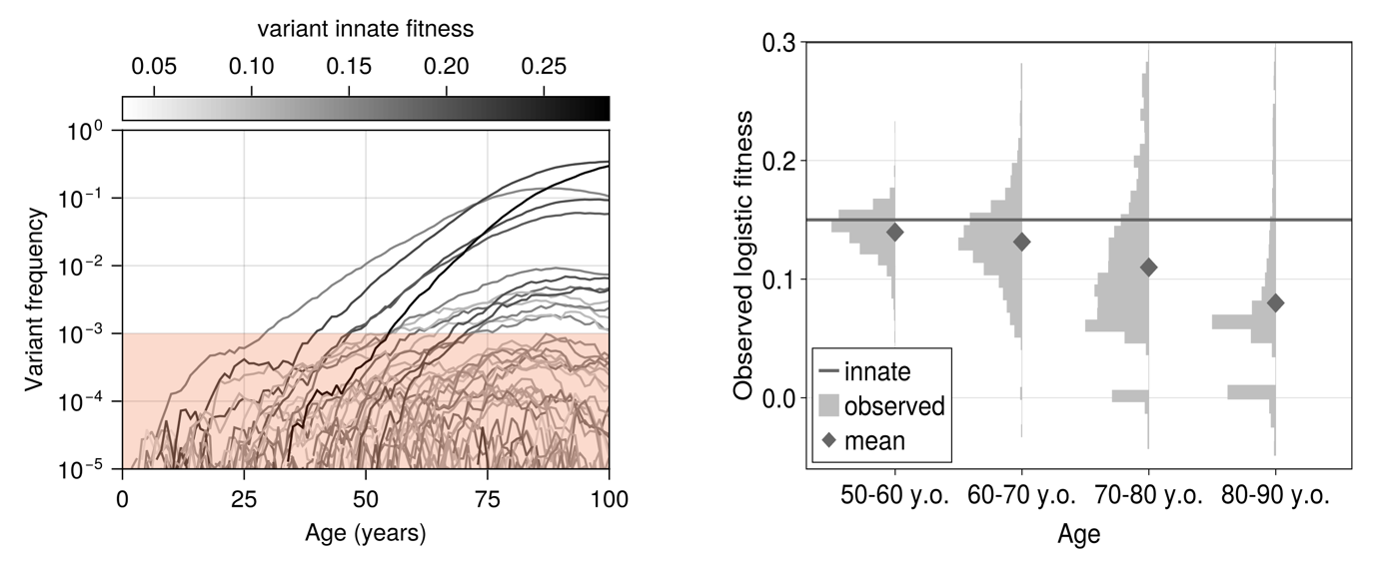
Given these results, we next asked ourselves how else we might observe clonal competition if it occurred. In particular, we wondered if one could measure it from changes in HSC genetic heterogeneity? To answer this question, we modeled the expected change of the site frequency spectrum (SFS) – the distribution (i.e. histogram) of variant frequencies in a population – in HSCs from newborns to octogenarians.
The dynamics of the SFS are complex. Low frequency variants are distributed deterministically (following power laws) – because there are many of them – but high frequency variants – being rare – present highly stochastic behavior. From our model we predicted two transitions: a first change from development to homeostasis, and a second from homeostasis to observable clonal growth. Both would imply measurable changes of the distributions of low and high frequency variants. When inspecting single-cell HSC data from Mitchell et al. [2], this is exactly what we observed!
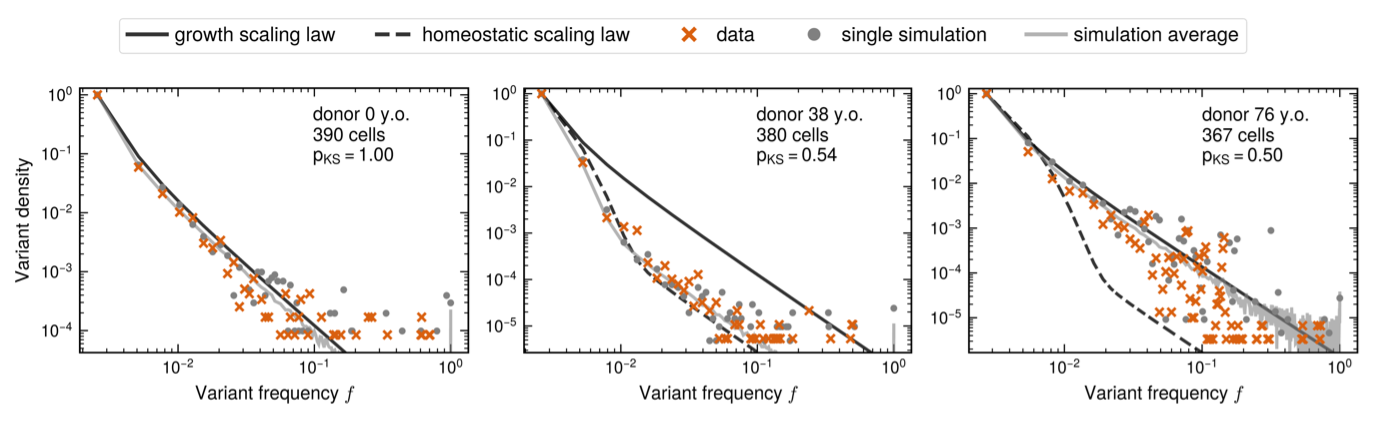
So, could we go beyond a qualitative description and derive quantitative predictions for the fitness distribution and arrival rate of clones? To tackle this question, we developed Approximate Bayesian Inference (ABC) frameworks for trajectory and SFS data independently. To perform ABC, we needed summary statistics to compare data and model. For clone trajectories, we used the distribution of clone sizes and the distribution of trajectory-fitted logistic fitnesses. For the SFS dynamics with age, we used the number of detectable clones at 1% frequency and the best fit to 7 out of 8 data points of the SFS.
From these statistics we used the ABC to estimate posterior distributions for the mean and standard deviation of the CH fitness distribution and the clone arrival rate per year. These independent inferences – one on clone trajectories and the other on the SFS dynamics – arrived at nearly identical results! This suggests to us that clonal competition can provide a meaningful and consistent explanation for observed complex dynamics in CH.
Finally, we’d like to applaud everyone on the initial papers, both for their incredible work and for making all the data available and accessible. Ultimately this growing pool of outstanding and publicly available data is what allows us as a community to dig deeper into the fascinating clonal dynamics of haematopoiesis and work together to improve our understanding of somatic evolution!
References
- Mon Père N, Terenzi F, Werner B. 2024. ‘The dynamic fitness landscape of ageing haematopoiesis through clonal competition’. bioRxiv, doi:10.1101/2024.04.16.589764.
- Mitchell, Emily, Michael Spencer Chapman, Nicholas Williams, Kevin J. Dawson, Nicole Mende, Emily F. Calderbank, Hyunchul Jung, et al. 2022. ‘Clonal Dynamics ofHaematopoiesis across the Human Lifespan’. Nature, June, 1–8. doi:10.1038/s41586-022-04786-y.
- Fabre, Margarete A., José Guilherme de Almeida, Edoardo Fiorillo, Emily Mitchell, Aristi Damaskou, Justyna Rak, Valeria Orrù, et al. 2022. ‘The Longitudinal Dynamics and Natural History of Clonal Haematopoiesis’. Nature, June, 1–8. doi:10.1038/s41586-022-04785-z.
- Zeventer, Isabelle A. van, Aniek O. de Graaf, Jonas B. Salzbrunn, Ilja M. Nolte, Priscilla Kamphuis, Avinash Dinmohamed, Bert A. van der Reijden, Jan Jacob Schuringa, Joop H. Jansen, and Gerwin Huls. 2023. ‘Evolutionary Landscape of Clonal Hematopoiesis in 3,359 Individuals from the General Population’. Cancer Cell 41 (6): 1017-1031.e4. doi:10.1016/j.ccell.2023.04.006.
© 2026 - The Mathematical Oncology Blog