The context-dependent cost of high ploidy
Opposing selective forces shape tumor ploidy
Tipping cancer cells over the edge: the context-dependent cost of high ploidy
Noemi Andor, Philipp M Altrock, Navami Jain, Ana P Gomes
Read the paper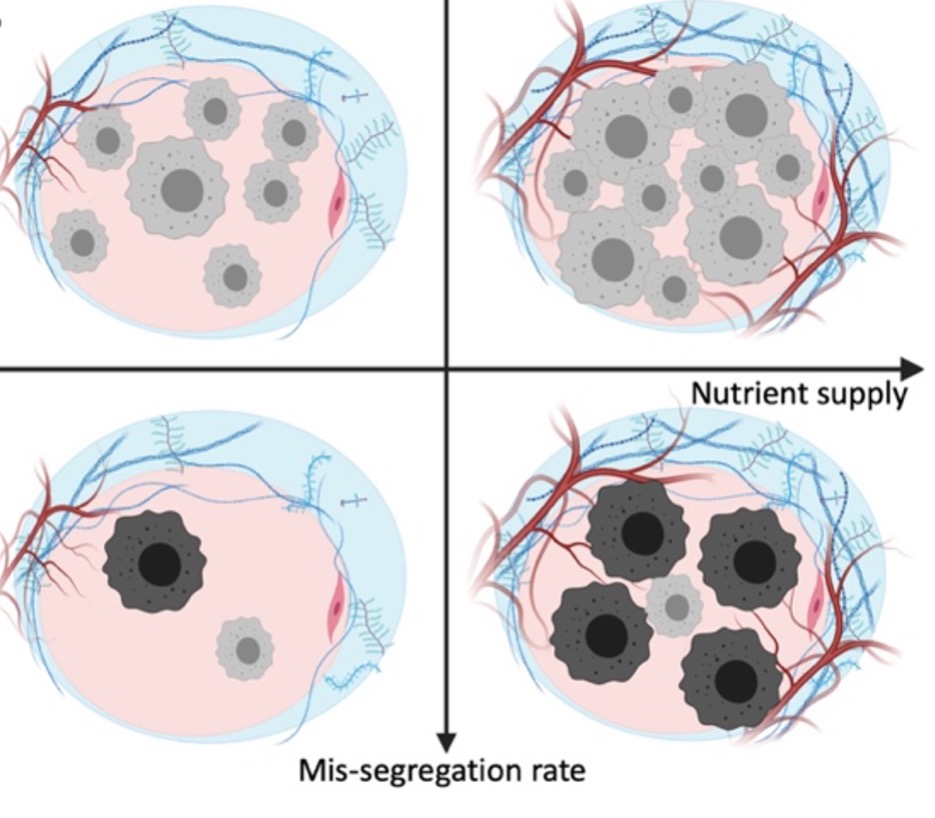
Figure 1: Opposing selective forces shape tumor ploidy. We distinguish cells along two dimensions: their load of somatic copy number alterations (SCNAs; grayscale) and their ploidy (drawn as cell size). Compared to low ploidy cells, high ploidy cells are more likely to survive mutations and accumulate high SCNA loads (y-axis). High ploidy cells require more nutrients for growth (x-axis), setting high ploidy cells at a disadvantage when competing in nutrient-scarce micro-environments.
Raw materials needed to generate deoxynucleotides, the building blocks of DNA, are candidate rate-limiting factors for the evolution of high-ploidy cancer cells. Specifically, we discuss Oxygen, Phosphate and Glucose as limiting substrates of dNTP synthesis in gastric and brain cancers: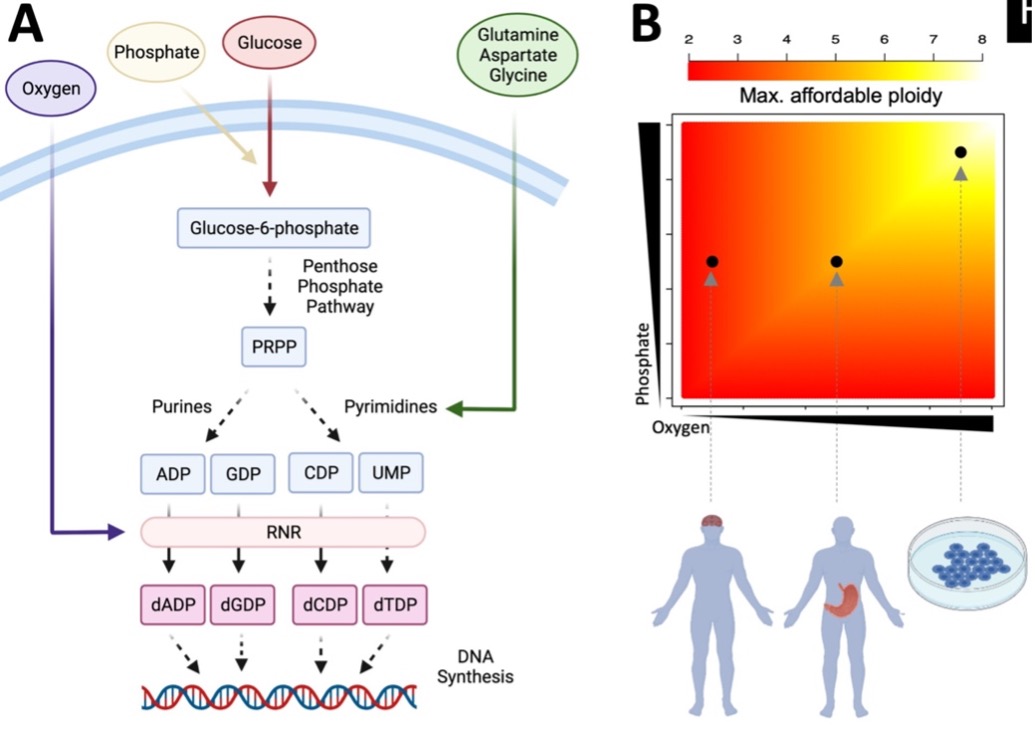 For example, Oxygen levels lie at the intersection of several phenomena that are unique to Glioblastoma (GBM) – an aggressive type of brain cancer. Stark gender differences in GBM pathology correlate with the sexual dimorphism in physiologic oxygen perfusion levels in the brain4,5. Pseudopalisading necrosis in GMB co-occurs near steep oxygen gradients6. Although oxygen saturation does not fall below 4% in most tissue types, in brain tissue oxygen levels often go down to 0.5%7. These studies support the idea that triploid and tetraploid states can often be a disadvantage in the brain, as they demand dNTP synthesis rates that cancer cells cannot sustain due to inherent low abundance of O2 in brain tissue. Without the capacity to sustain double the amount of DNA, these tumors are limited in their evolvability and may be trapped at local fitness optima.
While we focused on gastro-intestinal and brain environments, the hypothesis could also apply to several other cancer types and may explain the variable ploidy levels seen across different primary and metastatic sites:
For example, Oxygen levels lie at the intersection of several phenomena that are unique to Glioblastoma (GBM) – an aggressive type of brain cancer. Stark gender differences in GBM pathology correlate with the sexual dimorphism in physiologic oxygen perfusion levels in the brain4,5. Pseudopalisading necrosis in GMB co-occurs near steep oxygen gradients6. Although oxygen saturation does not fall below 4% in most tissue types, in brain tissue oxygen levels often go down to 0.5%7. These studies support the idea that triploid and tetraploid states can often be a disadvantage in the brain, as they demand dNTP synthesis rates that cancer cells cannot sustain due to inherent low abundance of O2 in brain tissue. Without the capacity to sustain double the amount of DNA, these tumors are limited in their evolvability and may be trapped at local fitness optima.
While we focused on gastro-intestinal and brain environments, the hypothesis could also apply to several other cancer types and may explain the variable ploidy levels seen across different primary and metastatic sites:
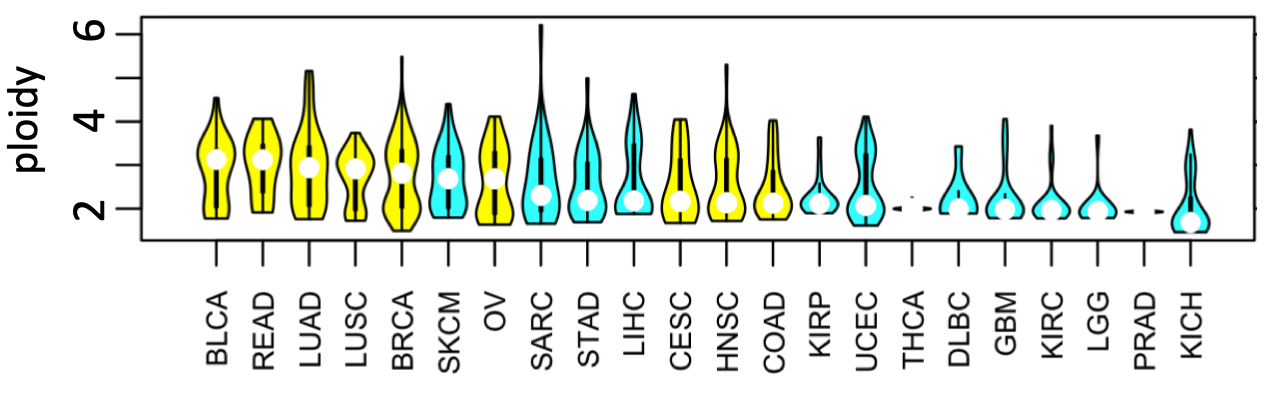 If this hypothesis is true, the implications are broad. It would explain vast differences in the extent of inter-tumor karyotype heterogeneity. For example, the vast majority of GBM fall into a single, well-defined karyotype defined by chromosome 7 amplification and chromosome 10 deletion, whereas gastric cancers have highly variable karyotypes. It may contribute to explain why agents that block dNTP production8 work well in combination with DNA damaging agents -- because they complement each other in targeting high- and low-ploidy cells that co-exist in a tumor.
If a tetraploid subclone expands, it consistently does so early in tumor evolution9,10, when cell density is still low and competition for nutrients is comparatively weak, suggesting these conditions favor the growth of tetraploid cells. We recently developed a mathematical model of co-evolving high and low ploidy cells11,12, accounting for the increased resource demands of high-ploidy cells. Models such as this could be used to evaluate competition between low and high ploidy cells during early stages of tumor evolution. Understanding the resource cost of high ploidy can help uncover its therapeutic vulnerabilities across tissue sites with versatile energy supplies.
If this hypothesis is true, the implications are broad. It would explain vast differences in the extent of inter-tumor karyotype heterogeneity. For example, the vast majority of GBM fall into a single, well-defined karyotype defined by chromosome 7 amplification and chromosome 10 deletion, whereas gastric cancers have highly variable karyotypes. It may contribute to explain why agents that block dNTP production8 work well in combination with DNA damaging agents -- because they complement each other in targeting high- and low-ploidy cells that co-exist in a tumor.
If a tetraploid subclone expands, it consistently does so early in tumor evolution9,10, when cell density is still low and competition for nutrients is comparatively weak, suggesting these conditions favor the growth of tetraploid cells. We recently developed a mathematical model of co-evolving high and low ploidy cells11,12, accounting for the increased resource demands of high-ploidy cells. Models such as this could be used to evaluate competition between low and high ploidy cells during early stages of tumor evolution. Understanding the resource cost of high ploidy can help uncover its therapeutic vulnerabilities across tissue sites with versatile energy supplies.
References
- Campbell PJ, Getz G, Stuart JM, Jennings JL, Stein LD, Perry MD, et al. Pan-cancer analysis of whole genomes. Nature. 2020;578:82–93.
- López S, Lim EL, Horswell S, Haase K, Huebner A, Dietzen M, et al. Interplay between whole-genome doubling and the accumulation of deleterious alterations in cancer evolution. Nat Genet. 2020;52:283–93.
- Andor, Noemi, Altrock, Philipp, Jain, Navami, Gomes, Ana. Tipping cancer cells over the edge: the context-dependent cost of high ploidy. Cancer Res Press. 2021;
- Parkes LM, Rashid W, Chard DT, Tofts PS. Normal cerebral perfusion measurements using arterial spin labeling: reproducibility, stability, and age and gender effects. Magn Reson Med. 2004;51:736–43.
- Amen DG, Trujillo M, Keator D, Taylor DV, Willeumier K, Meysami S, et al. Gender-Based Cerebral Perfusion Differences in 46,034 Functional Neuroimaging Scans. J Alzheimers Dis. 2017;60:605–14.
- Hambardzumyan D, Bergers G. Glioblastoma: Defining Tumor Niches. Trends Cancer. 2015;1:252–65.
- Haque N, Rahman MT, Abu Kasim NH, Alabsi AM. Hypoxic culture conditions as a solution for mesenchymal stem cell based regenerative therapy. ScientificWorldJournal. 2013;2013:632972.
- Pyrimidine Analogues. LiverTox Clin Res Inf Drug-Induc Liver Inj [Internet]. Bethesda (MD): National Institute of Diabetes and Digestive and Kidney Diseases; 2012 [cited 2021 Mar 21]. Available from: http://www.ncbi.nlm.nih.gov/books/NBK548604/
- Bielski CM, Zehir A, Penson AV, Donoghue MTA, Chatila W, Armenia J, et al. Genome doubling shapes the evolution and prognosis of advanced cancers. Nat Genet. 2018;50:1189–95.
- Dewhurst SM, McGranahan N, Burrell RA, Rowan AJ, Grönroos E, Endesfelder D, et al. Tolerance of whole-genome doubling propagates chromosomal instability and accelerates cancer genome evolution. Cancer Discov. 2014;4:175–85.
- Kimmel GJ, Dane M, Heiser LM, Altrock PM, Andor N. Integrating mathematical modeling with high throughput imaging explains how polyploid populations behave in nutrient-sparse environments. Cancer Res [Internet]. 2020 [cited 2020 Oct 5]; Available from: https://cancerres.aacrjournals.org/content/early/2020/09/16/0008-5472.CAN-20-1231
- Andor, N., Balanced phenotypic trade-offs in polyploid cancer populations slow down invasion; The Mathematical Oncology Blog. 2020. Available from: https://mathematical-oncology.org/blog/trade-offs-polyploid-cancer.html
© 2026 - The Mathematical Oncology Blog