Mistic, an open-source multiplexed image t-SNE
Behind the paper
Mistic, an open-source multiplexed image t-SNE
Sandhya Prabhakaran, Chandler Gatenbee, Mark Robertson-Tessi, Jeffrey West, Amer A. Beg, Jhanelle Gray, Scott Antonia, Robert A. Gatenby, Alexander R. A. Anderson
Read the preprint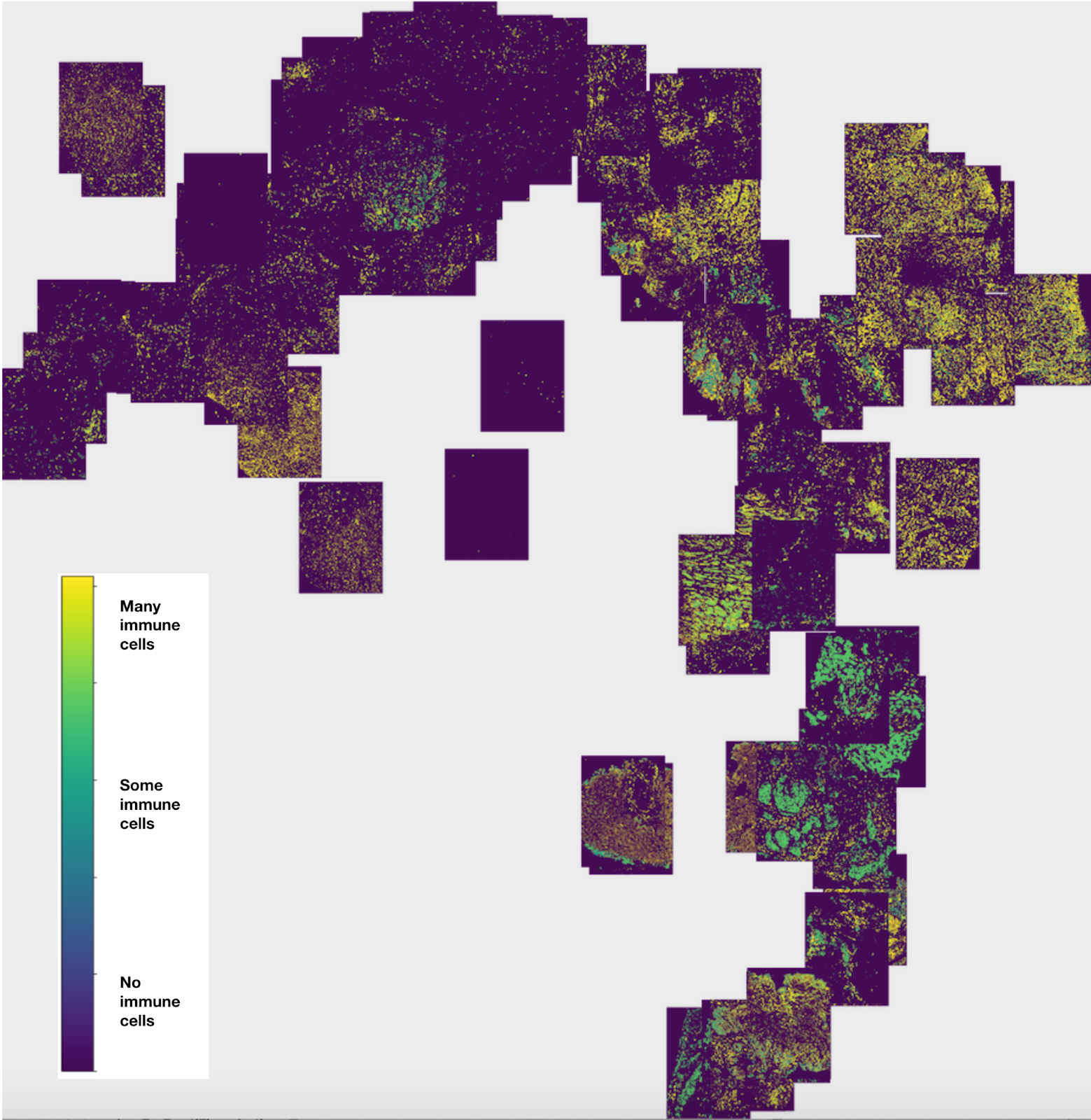
Figure 1: Image t-SNE showing immune cell gradient across multiple multiplexed images
We developed Mistic that generates such an image t-SNE. Mistic’s GUI is shown in Figure 2. Mistic generates thumbnails for images based on user preferences and arranges them in different layouts: (left) user-predefined coordinates (e.g. from t-SNE or UMAP analysis), (middle) using a grid layout, or (right) random coordinates (Figure 3).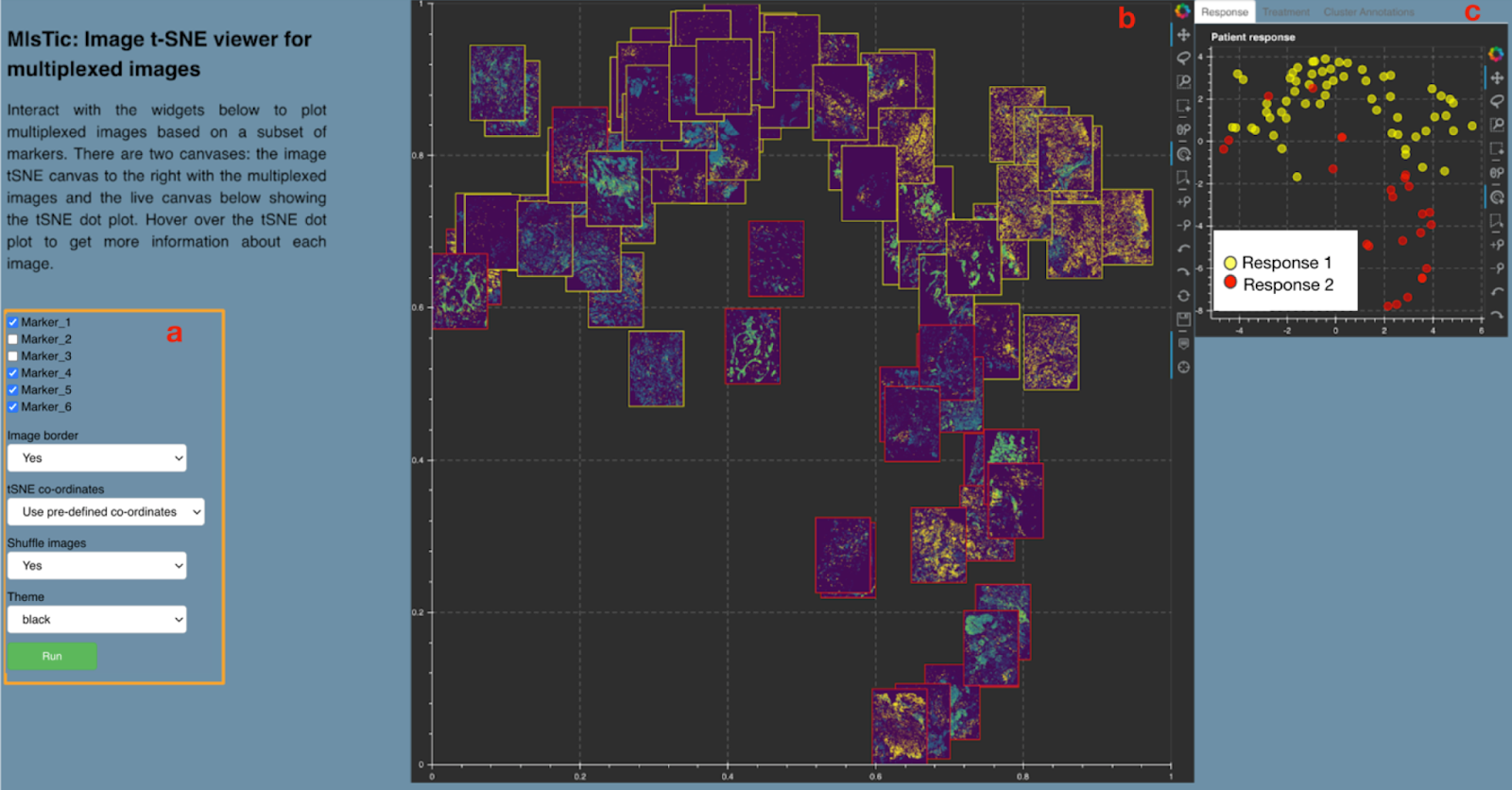
Figure 2: Mistic GUI: a. User-input panel where markers can be selected, new or pre-defined image display coordinates can be chosen, etc. b. Static canvas showing the image t-SNE colored and arranged as per user inputs. c. Live canvas showing the corresponding t-SNE scatter colored based on metadata.
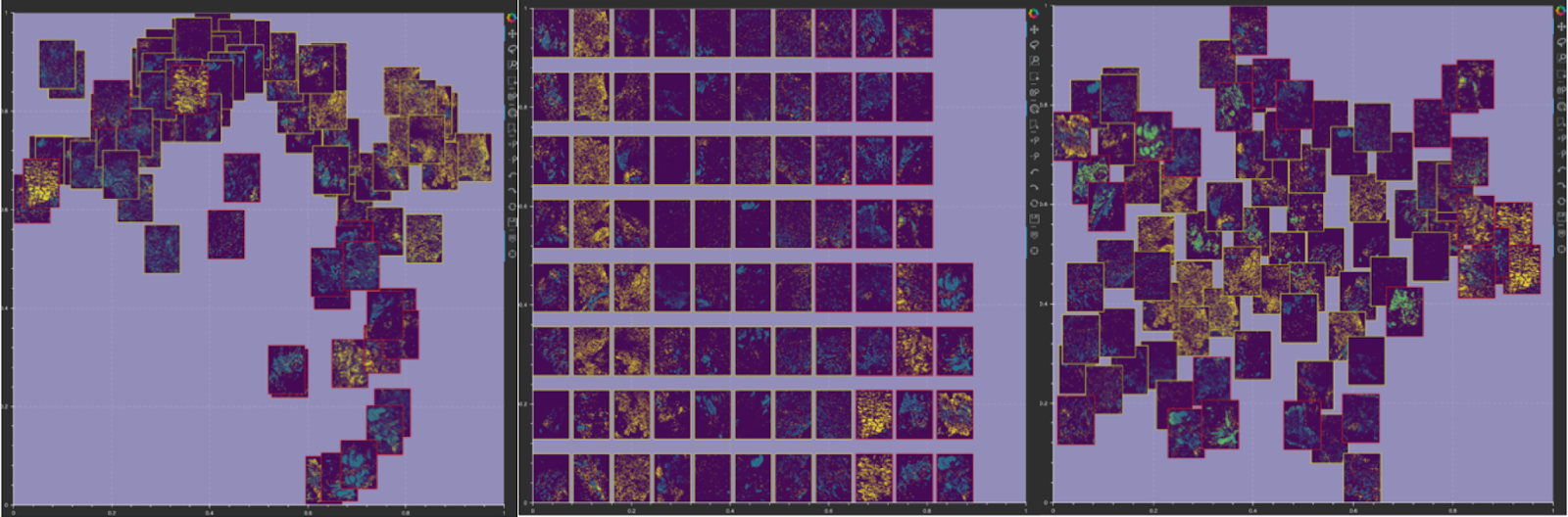
Figure 3: Image layout in Mistic’s static canvas: (i) based on user-defined t-SNE coordinates (left); (ii) vertical rows (center); (iii) randomly placed (right).
We tested Mistic on multiplexed images from Perkinelmer Vectra. Mistic also allows metadata of the multiplexed images to be displayed. Currently there is no freely available tool to generate such image t-SNEs. Mistic is developed using Bokeh and Python, and can be downloaded directly from the IMO GitHub pages at: https://github.com/MathOnco/Mistic. A preprint with further details on how to use Mistic is available on BioRxiv here. Future work involves experimenting Mistic with higher dimensional images and images generated using different technologies.© 2026 - The Mathematical Oncology Blog