Integrating Quantitative Assays with Biologically Based Mathematical Modeling for Predictive Oncology
Anum S. Kazerouni, Manasa Gadde, Andrea Gardner, David A. Hormuth II, Angela M. Jarrett, Kaitlyn E. Johnson, Ernesto A.B. F. Lima, Guillermo Lorenzo, Caleb Phillips, Amy Brock, Thomas E. Yankeelov
Read the publication
The complexity of cancer cannot be understated. Decades of cancer research studying the dynamics of protein expression, cell populations, and tissue microenvironments have made it evident that a tumor’s evolution is governed by numerous spatially and temporally heterogeneous processes. Despite dramatic increases in our knowledge of cancer biology across scales, we currently lack an understanding of the governing principles of a tumor’s progression and response to treatment. Seeking to address this gap in understanding, researchers within the field of mathematical oncology have worked to develop mathematical models based on the physiological processes of cancer. With direct incorporation of biological mechanisms into their formalism, these models can be used to simulate cancer phenomena and forecast cancer progression and response.
This sentiment is not new, as models of cancer dynamics go back over half a century
1. However, historically, these models have been designed in terms of parameters or variables that are difficult to measure experimentally, thereby limiting their practicality and ability to be applied in a clinical setting. This yields a state of the field with many mathematical models of varying complexities but few that have been systematically and rigorously tested with experimental data. It is vital to recognize the importance of framing mathematical models with measurable parameters; i.e., designing models that allow incorporation of experimental or clinical data
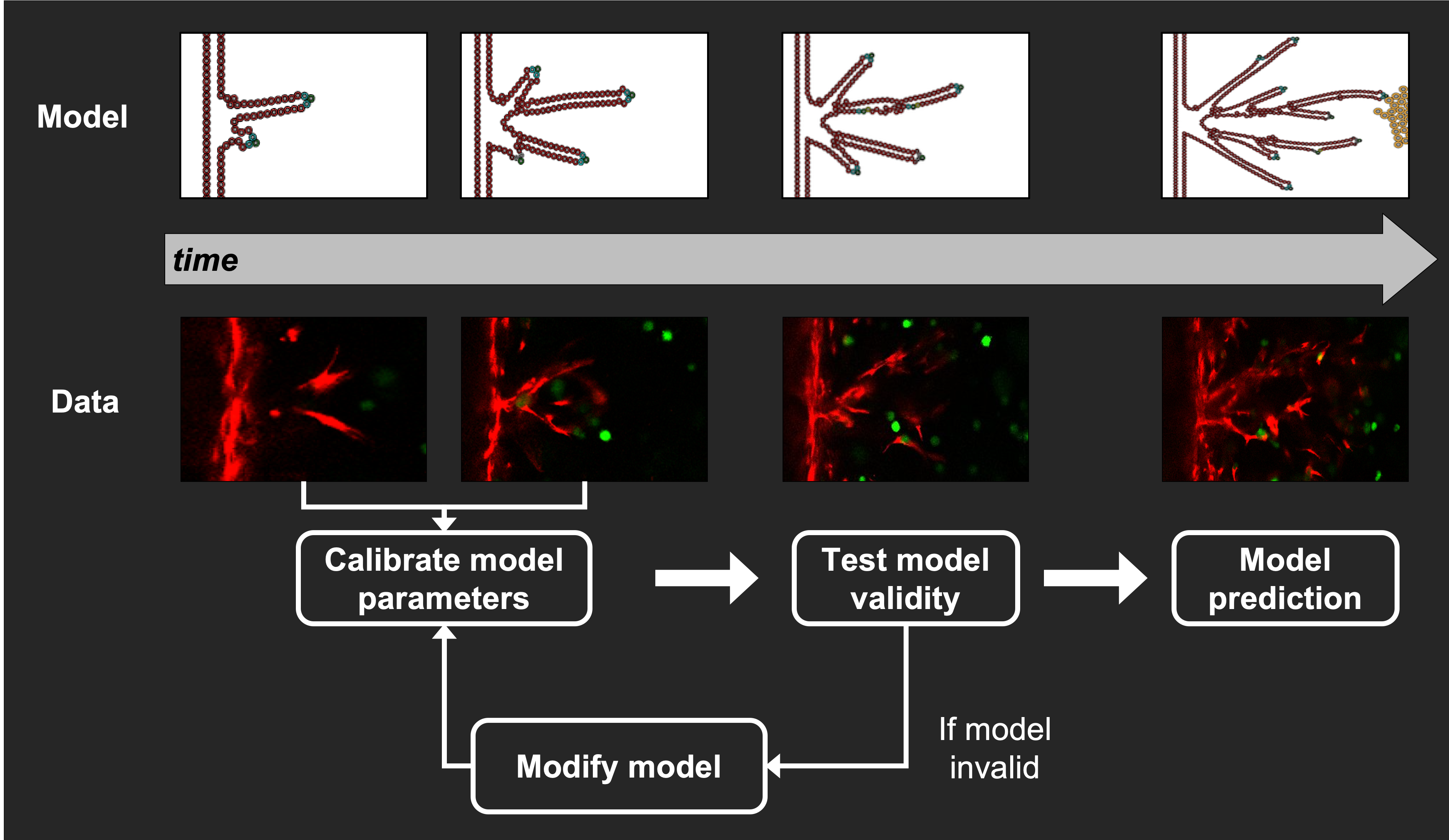
2,3. Without measurable parameters, we are left with challenging problems in ensuring parameter identifiability and model calibration. Additionally, the integration of mathematical models with experimental data facilitates their use in practical settings as they are constrained by credible equations and parameters. Simply, if we are to predict the future, we must be able to see the past (constrained and validated our model with many data sets) and understand the present (initialized the model with the current state of the system).
Recent advances in experimental assays and imaging techniques have enabled quantitative measurement of cancer dynamics that can be used to parameterize models of cancer growth and response. Such data-fortified models can be used to guide experimental design and focus the selection of biological measurements to be acquired. These models can also lend insight into cancer mechanisms that may be difficult to measure experimentally. Furthermore, integration of quantitative measurements into mathematical models of cancer can allow for patient-specific modeling (where a model is parameterized by measurements from an individual patient), enabling in silico investigation of dose regimens, treatment schedules, the influence of certain biological mechanisms of interest, etc.
In this review, we discuss biological assays that provide quantitative measurement of tumor characteristics at the cellular, microenvironmental, tissue, and organ scales. We describe how data from these assays can then be incorporated into mechanism-based models for the improved prediction of tumor growth and treatment response. A critical component towards effective mathematical oncology research is close collaboration between experimentalists and mathematicians to iterate through experimental design and model development together. This review seeks to expose both parties to opportunities for data integration into mathematical models of cancer, efforts that can accelerate research towards a comprehensive theory of cancer.
References
- Araujo, R. A history of the study of solid tumour growth: the contribution of mathematical modelling. Bulletin of Mathematical Biology 66, 1039–1091 (2004).
- Yankeelov, T. E. et al. Clinically Relevant Modeling of Tumor Growth and Treatment Response. Science Translational Medicine 5, 187ps9-187ps9 (2013).
- Yankeelov, T. E., Quaranta, V., Evans, K. J. & Rericha, E. C. Toward a Science of Tumor Forecasting for Clinical Oncology. Cancer Research 75, 918–923 (2015).