Defining metaphenotypes
Behind the Paper
Tumor-immune metaphenotypes orchestrate an evolutionary bottleneck that promotes metabolic transformation
Jeffrey West, Frederika Rentzeperis, Casey Adam, Rafael Bravo, Kimberly A. Luddy, Mark Robertson-Tessi, Alexander R. A. Anderson
Read the preprintThe metaphenotype
In this blog post, I review a new concept underlying our most recent preprint: the concept of the "meta-phenotype." The preprint illustrates an agent-based model of cancer metabolism (built on previous work in Sandy Anderson's lab, published in Cancer Research1). The model consists of two phenotypes: Acid Resistant and Glycolysis. Highly glycolytic cells produce acid, in turn leading to acid-mediated invasion of tumor into normal tissue. This process of acid-mediated invasion can be classified as a "collective cellular phenomena" - not driven by a single cell but by a collective action of multiple cells working in concert. An example is shown below, where I've seeded a simulation with a low Glycolysis phenotype (blue) and a high Glycolysis phenotype (purple). When seeded randomly, invasion does not occur (top row), because the volumetric acid produced by glycolytic cells is not enough to cause invasion. But when purple, high Glycolysis cells are arbitrarily seeded on the rim, invasion occurs due to the collective acidification of cells within close proximity.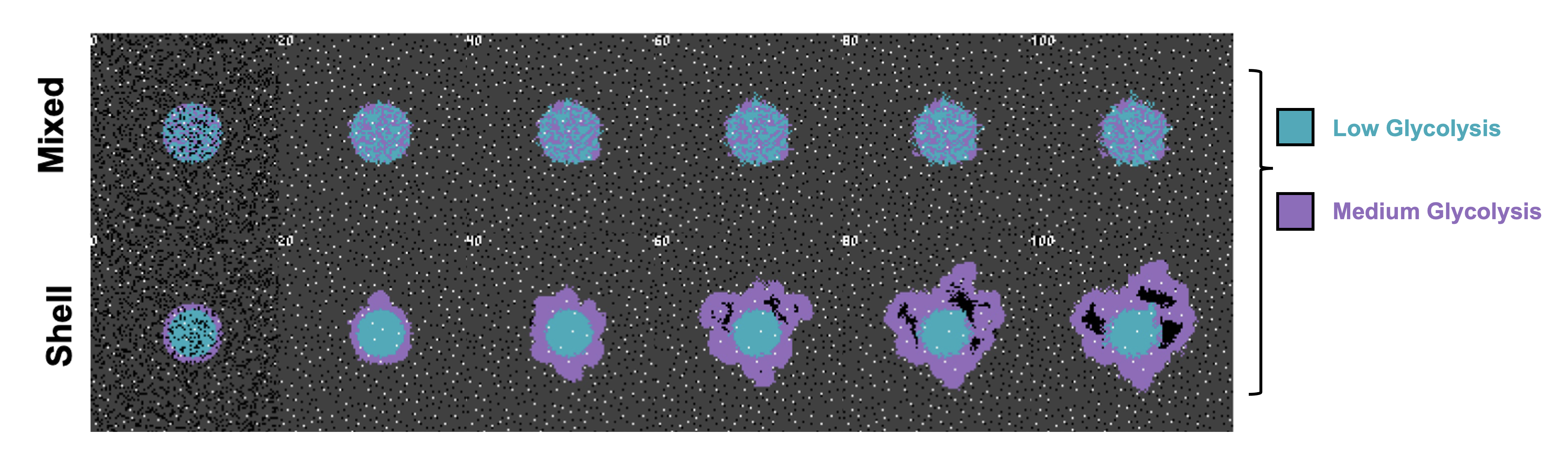
Figure 1: A contrived example illustrating how the initial configuration of heterogeneity leads to differential outcomes due to group selection dynamics. A low glycolysis phenotype (blue) and a high glycolysis phenotype (purple) compete for resources. Top row: a mixed configuration leads to no evolution. Acid-mediated invasion does not occur because the volumetric acid produced by aggressive cells is not enough to cause invasion when highly glycolytic cells are seeded far apart. Bottom row: In contrast, artificially placing the aggressive high glycolysis phenotypes on the rim leads to invasion from increased volumetric acid via a group-effect.
Do we really need to redefine the phenotype?
We extended this baseline model of cancer metabolism to include immune predation: T-cells that target tumor cells. We've also added a third phenotype: Glycolysis, Acid Resistant, and now PD-L1. By definition, immune escape is a collective cellular phenomenon (requiring at least two cells: a tumor and an immune cell). But this minor model extension allows for a wide range of additional collective behavior. In particular, we are interested in the collective phenotypes at play in immune escape. How do cells work together to avoid immune predation? Each cancer cell can have only three phenotypes (Glycolysis, Acid Resistant, PD-L1). But there are many more possible configurations of a cell's local context. The cellular context consists of neighboring cells and microenvironmental factors that can alter the effectiveness of each phenotype. For example, a cancer cell may have high PD-L1 expression and theoretically survive an immune interaction, but eventually die due to an acidic environment. The novelty here is that we define a new, higher-level cellular phenotype (a metaphenotype) which explicitly accounts for environmental factors and neighboring cell types. Sandy (along with Mark Robertson-Tessi) have been brainstorming this concept for a few years now:My own, shorthand definition of the metaphenotype: effective & collective. Effective: In our agent-based model, we can distinguish between cells who utilize a specific phenotype (effective) versus those who may use a phenotype (potential). For example, a cell might be PD-L1 positive, but not near a T-cell (potential immune escape, but not effective). Collective: As alluded before, we define meta-phenotypes that take into account 1) phenotype 2) cellular neighbors and 3) environmental effects. This allows for new, collective behaviors like mooching. A cell can rely on neighbors to provide a physical blockade against immune predation. A cell can live in an acidic niche (inactivating T-cells) created by neighboring cells.This emerged because the individual cellular phenotype was insufficient to define behaviors like acid mediated invasion. It might even be that, a specific range of phenotypic diversity is requirement to build a particular metaphenotype #ISEEC17
— Sandy Anderson (@ara_anderson) December 8, 2017
Immune Escape metaphenotypes
Figure 2 illustrates the concept of the metaphenotype. The Venn diagram shows the intersection of various phenotypes, providing our collective behavior. Here, we define 7 metaphenotypes that provide a mechanism of immune escape. The "null" metaphenotype is the lack of collective behavior: Immune Desert are cells which do not interact with T-cells. Next, we quantify two PD-L1 metaphenotypes: a counter-attack (tumor cell with high PD-L1 expression that has recently interacted with a T-cell; PD-L1 Attack, yellow), and a mooching PD-L1 (low PD-L1, with a high PD-L1 neighboring cell; Mooch PD-L1, blue). Two metaphenotypes rely on acid-inactivation: self-acidifying (highly glycolytic cells which secrete acid; Self-Acidify, pink) and non-producers (reside in acidic niche but do not produce acid; Mooch Acid, green). We also consider a proliferative phenotype that has recently divided into empty space (Proliferate Fast; red). Lastly, tumor cells also compete with immune cells for glucose (Starve Glucose; light blue). Importantly, each of these metaphenotypes (excluding Immune Desert) is contingent on a recent tumor- immune interaction, allowing us to track effective collective phenotypes: only metaphenotypes which survive an immune interaction.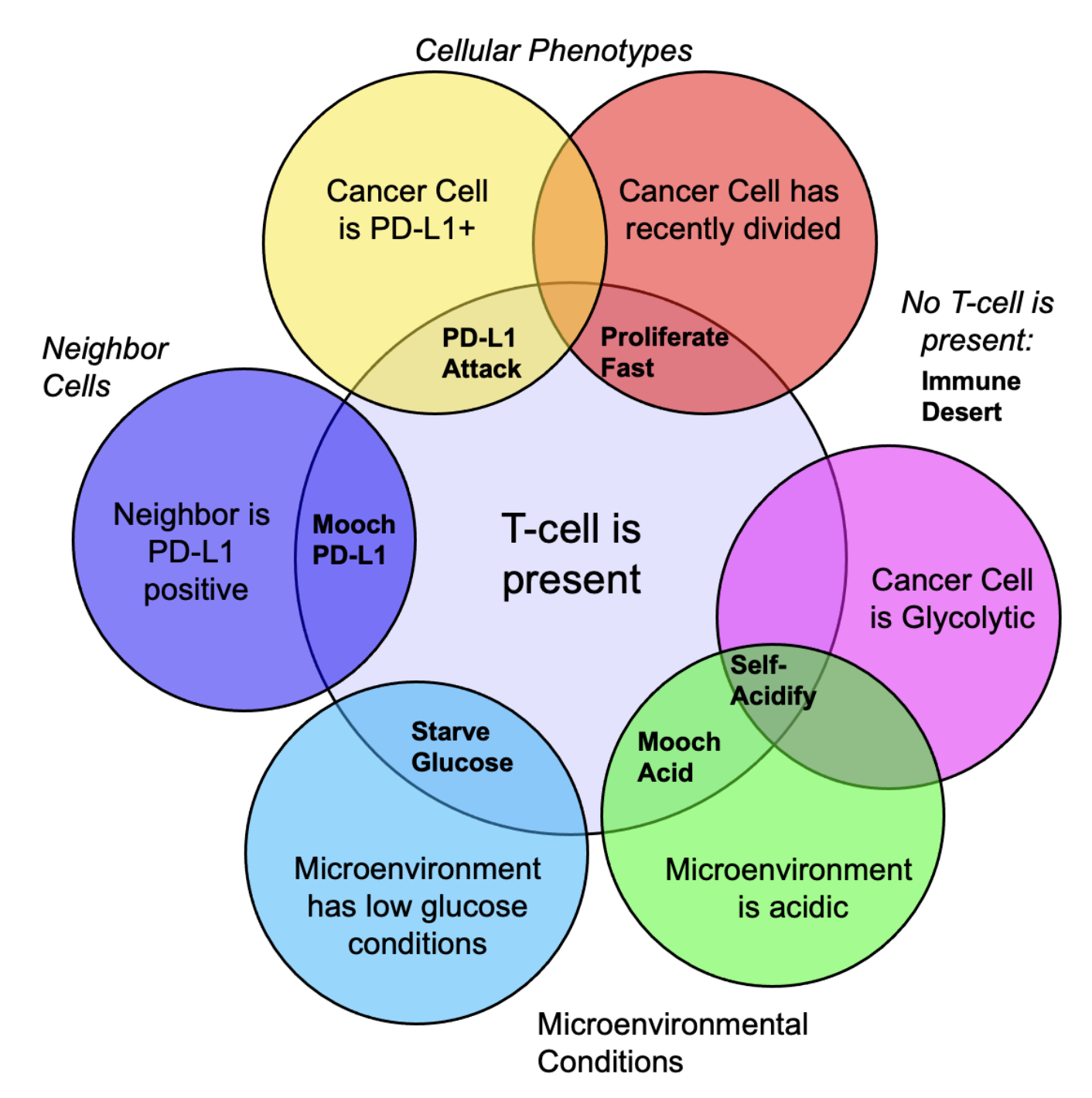
Figure 2: Defining metaphenotypes
The "lightbulb" moment for me came when I attempted to define all metaphenotypes on a scale from 0 to 1, where a value of 0 corresponds to a tumor cell with zero likelihood of surviving an immune interaction, and a value of 1 corresponds to a tumor cell with maximum likelihood of surviving the same immune targeting. For example, the "PD-L1 Attack" metaphenotype is defined as the probability of avoiding T-cell kill (based on PD-L1 expression), conditioned on a recent T-cell interaction $$ \texttt{PD-L1}(x,y)= \underbrace{(1-P_k)}_\text{Prob. avoiding T-cell kill} \times \underbrace{I(x,y)}_\text{recent T-cell interaction} $$ This insures that PD-L1 Attack is on the interval [0,1]. We repeat a similar process for all seven metaphenotypes and each cell has a vector of length 7 to define expression level for all 7: \(\vec{MP}=\{m_1,m_2, \dots, m_7\}\) where \(m_i \in [0,1]\).Conclusion
Precise characterization of collective phenotypes into metaphenotypes enables a straightforward explanation of the effect of treatment in a complex, multi-scale model. This characterization is necessary, in part, due to the fact that acid-mediated invasion is a collective phenotype phenomenon (figure 1). Immune escape is also, by definition, a collective phenomenon by requiring the presence of two cell types in close proximity: tumor and immune. Here, immune escape metaphenotypes are only defined in the context of recent immune-tumor interactions, by not using cellular phenotypes which are present but not actively employed (e.g. a PD-L1+ cell not in the immediate presence of an immune cell). Read the preprint to learn more details.References
- Robertson-Tessi, Mark, et al. "Impact of metabolic heterogeneity on tumor growth, invasion, and treatment outcomes." Cancer Research 75.8 (2015): 1567-1579.
© 2025 - The Mathematical Oncology Blog